Dimensionality Reduction & PCA
Lecture 32
Dr. Eric Friedlander
College of Idaho
CSCI 2025 - Winter 2026
Dimensionality
The Problem with High Dimensions
- Big Data can mean lots of observations (big \(n\)) or lots of variables (big \(p\)).
- Big \(p\) Problems:
- Visualization: Hard to plot > 3 dimensions.
- Curse of Dimensionality:
- Data becomes sparse.
- Distance metrics (Euclidean) lose meaning.
- Models can overfit.
Dimensionality Reduction
- Goal phrasing 1: Reduce the number of columns, while losing as little information as possible
- Goal phrasing 2: Extract lower-dimensional structure from our data
- Analogy: file compression
Which One is Compressed?


Which One is Compressed?


Idea
- We managed to:
- Reduce file size by 70%
- Not lose much information
- Extract underlying structure (a duck)
Thinking about structure in tabular data
Underlying Structure
- What dimension is our data in?
- What is the underlying structure here?
- What is the dimension of a plane?
Visualizing Plane
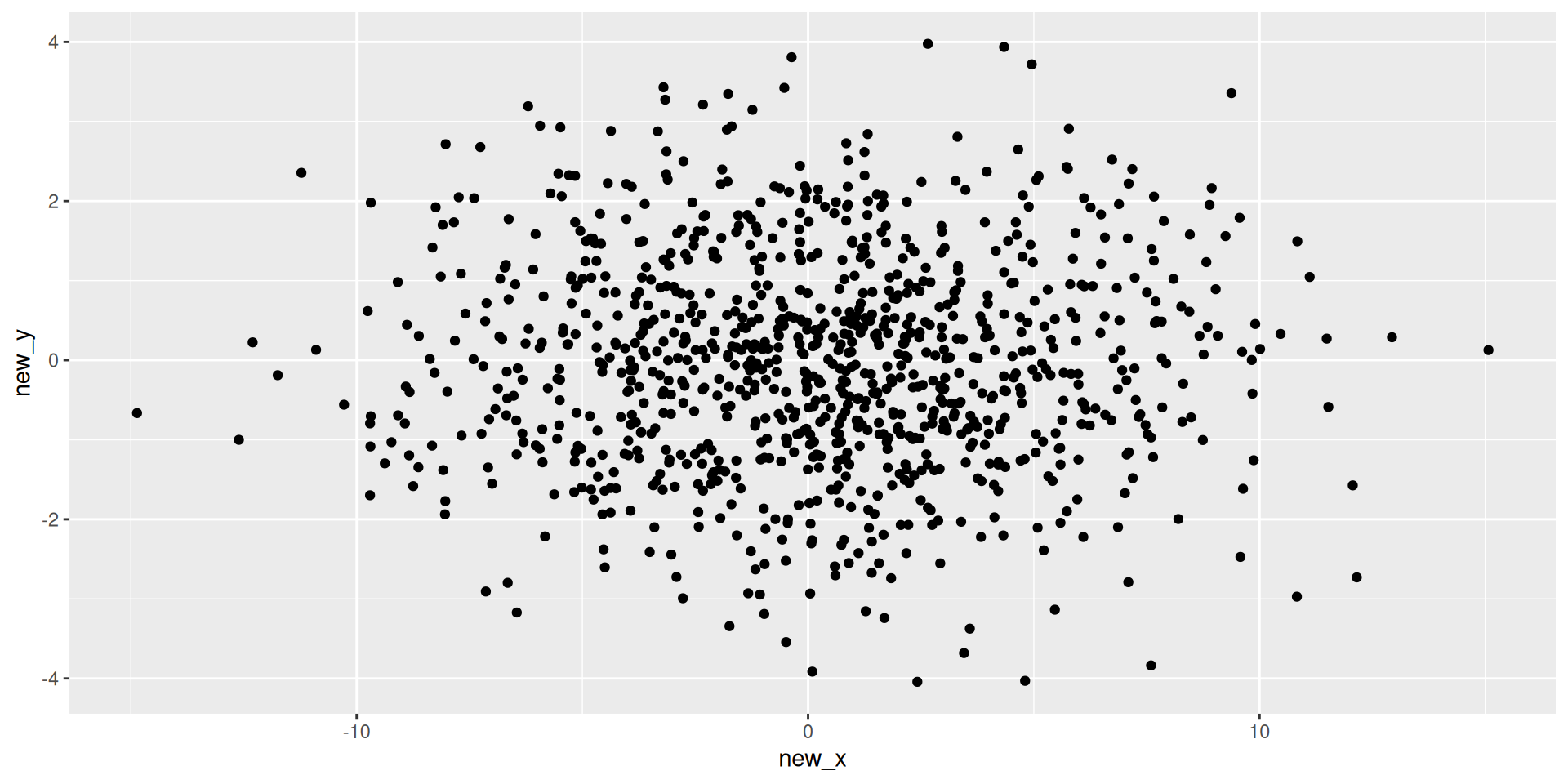
Thinking about structure in tabular data
Underlying Structure
- What dimension is our data in?
- What is the underlying structure here?
- What is the dimension of a line?
Visualizing 2D
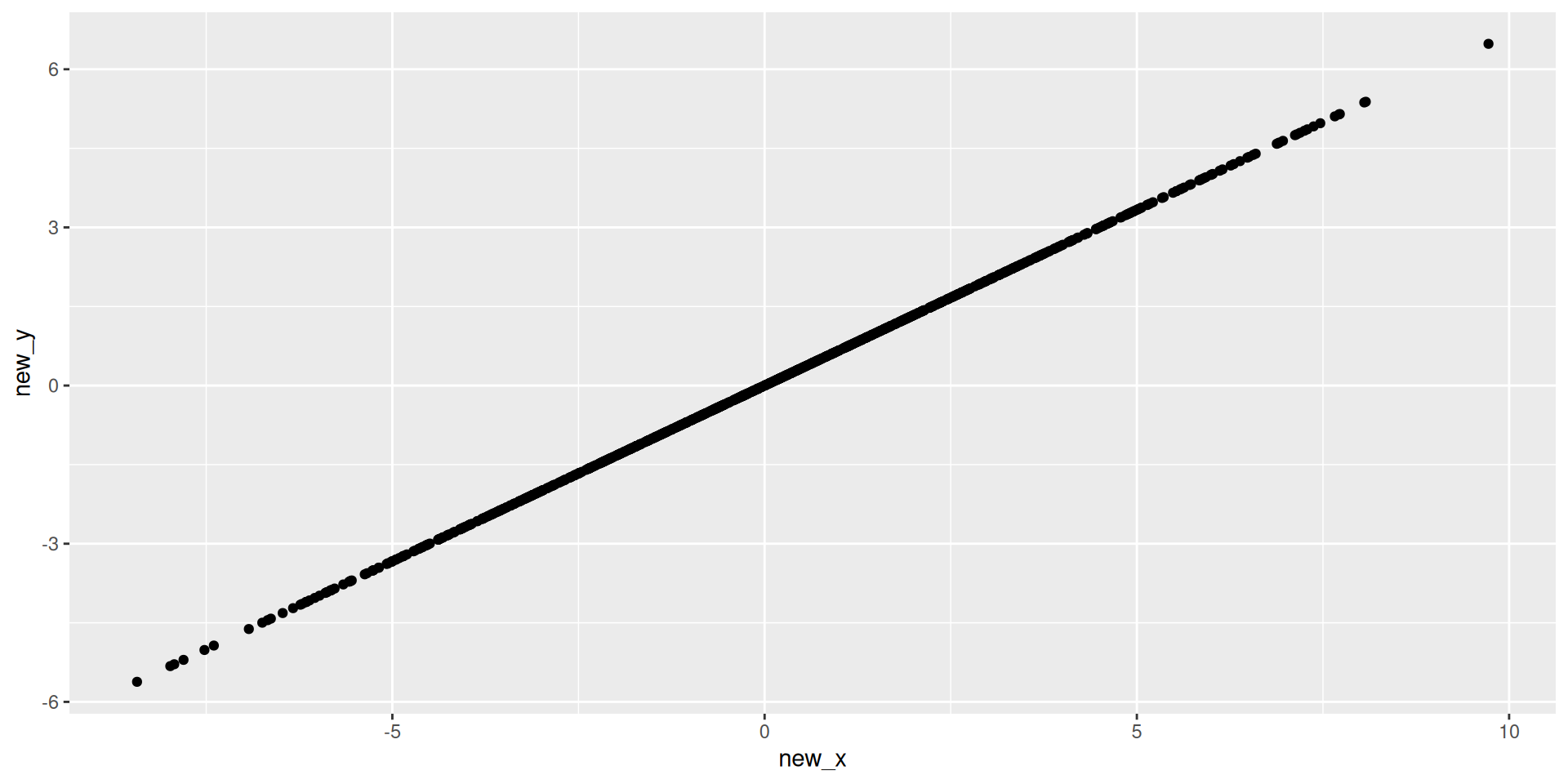
Visualizing 1D
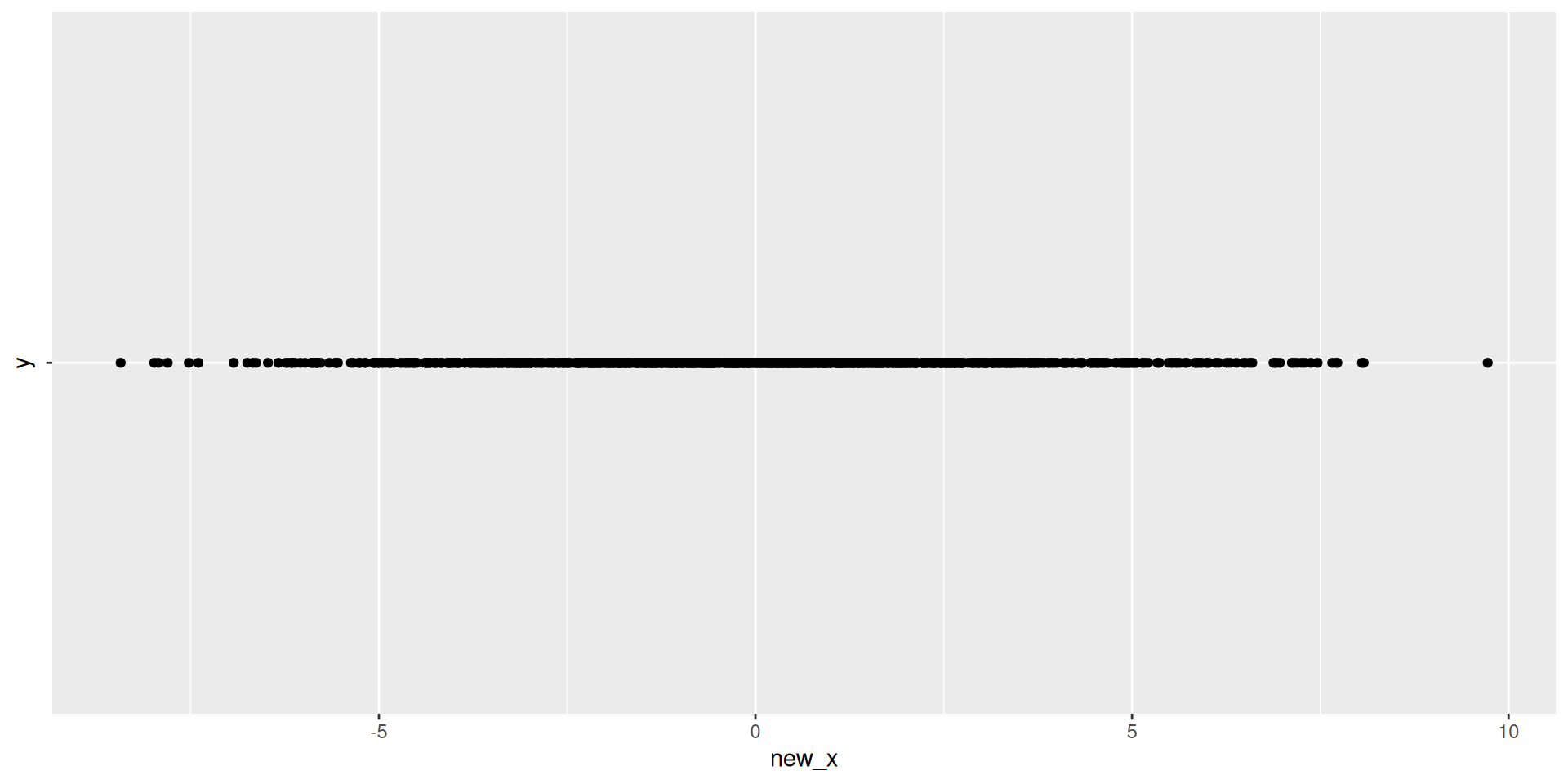
Visualizing 1D Data
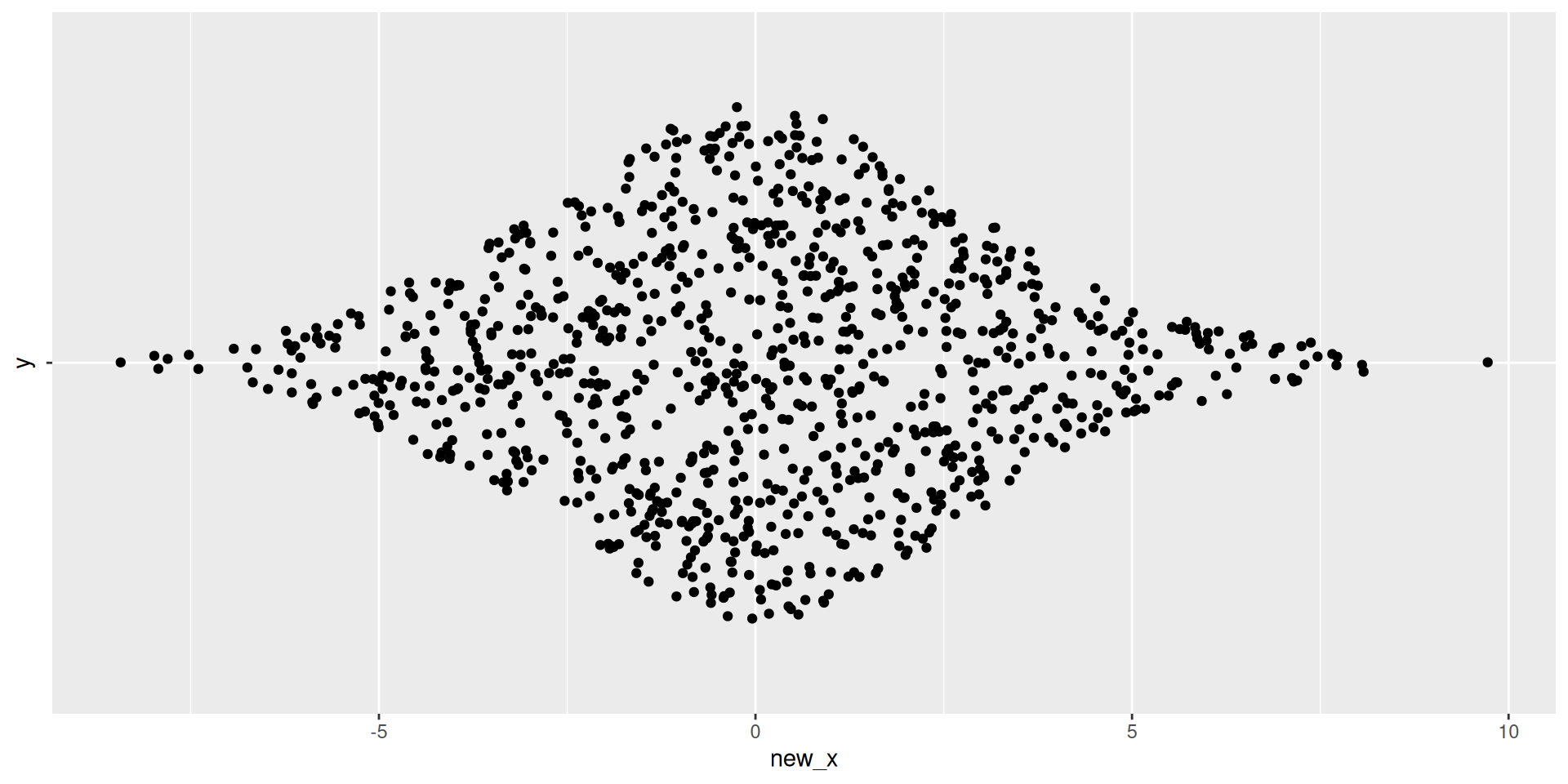
Thinking about structure in tabular data
Underlying Structure
- What dimension is our data in?
- What is the underlying structure here?
- What is the dimension of a plane?
Visualizing Plane
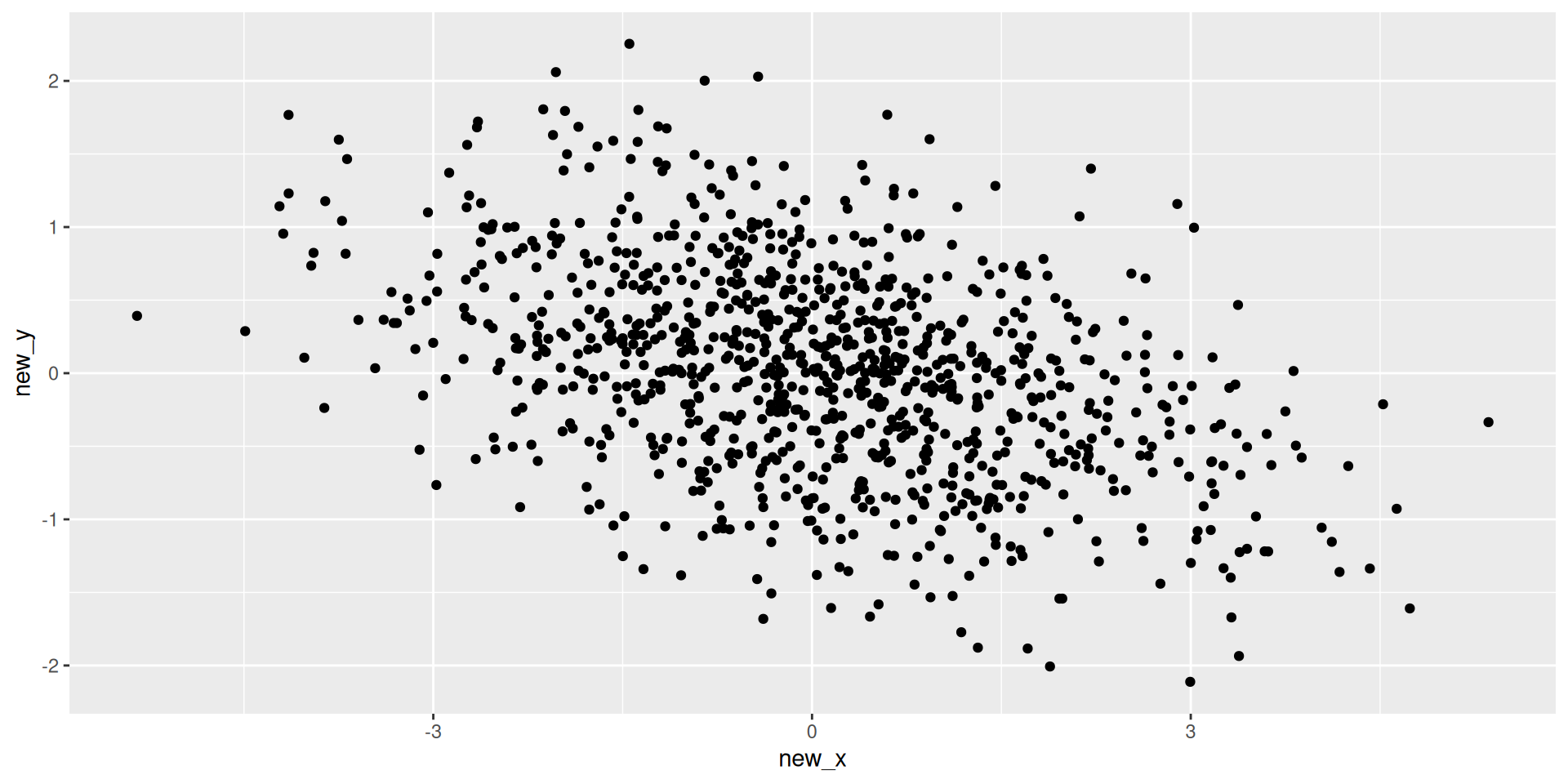
Discussion
- What’s the difference between the first two scenario’s and the third scenario?
- How much have we reduced the dimension?
- How much information have we lost?
Principal Component Analysis (PCA)
Vector’s and Projections
Basis Vectors and New Coordinates
- Plane above: \(z = x + y\)
- New Directions:
- New Direction 1: \(\vec{d}_1 = \langle 1, 1, 2\rangle\)
- New Direction 2: \(\vec{d}_2 = \langle 1, -1, 0\rangle\)
- New data:
- New \(x\): \(1\times x_{old} + 1\times y_{old} + 2\times z_{old}\)
- New \(y\): \(1\times x_{old} - 1\times y_{old} + 0\times z_{old}\)
- Note: Not quite correct, need to re-normalize
New Data
new_data <- data |>
mutate(new_x = x + y + 2*z,
new_y = x - y,
new_x = new_x/6, #re-normalizing
new_y = new_y/2)
new_data |> head() |> kable()| x | y | z | new_x | new_y |
|---|---|---|---|---|
| -0.5604756 | -0.9957987 | -1.5562744 | -0.7781372 | 0.2176615 |
| -0.2301775 | -1.0399550 | -1.2701325 | -0.6350663 | 0.4048888 |
| 1.5587083 | -0.0179802 | 1.5407281 | 0.7703640 | 0.7883443 |
| 0.0705084 | -0.1321751 | -0.0616667 | -0.0308334 | 0.1013418 |
| 0.1292877 | -2.5493428 | -2.4200550 | -1.2100275 | 1.3393153 |
| 1.7150650 | 1.0405735 | 2.7556384 | 1.3778192 | 0.3372458 |
What’s actually happening
- We are projecting each observation onto our new directions \(\vec{d}_1\) and \(\vec{d}_2\)
- Visualization
Projecting our data
Plotting these
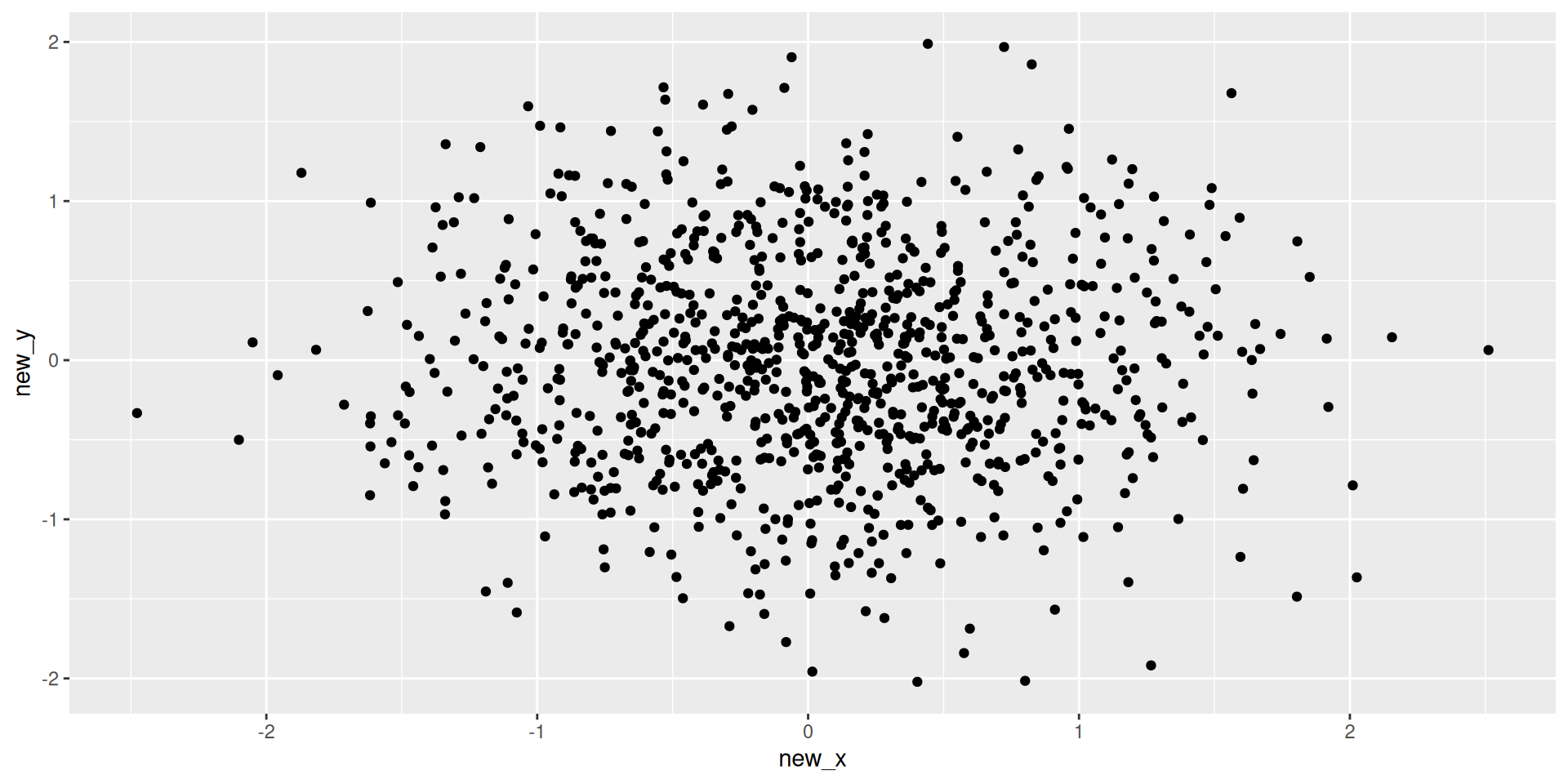
Principal Component Analysis (PCA)
PCA Vocabulary
- Principal Component (PC1): direction in \(p\)-dimensional space (e.g. \(\langle 1, 1, 2\rangle\))
- Scores: our new variables (e.g. \((-0.56\times 1 + -0.996\times 1 + -1.56\times 2)/6 = -0.778\))
- Loadings: For direction above’
- Loading on \(x\) is 1
- Loading on \(y\) is 1
- Loading on \(z\) is 2
Recall: Variance
- What is variance?
- Intuitively: what does variance measure?
- Variance: \(\frac{1}{n-1}\sum_{i=1}^n(x_i - \bar{x})^2\)
- Average of the squared distance from zero of each observation
Idea behind PCA
- Select first PC so variance of scores is the maximum
- Iteratively:
- Select next PC so variance of scores is maximize AND new PC is orthogonal to all other PCs
- What does orthogonal mean?
Easy Example
- Exercise: What should the first and second PCs be?
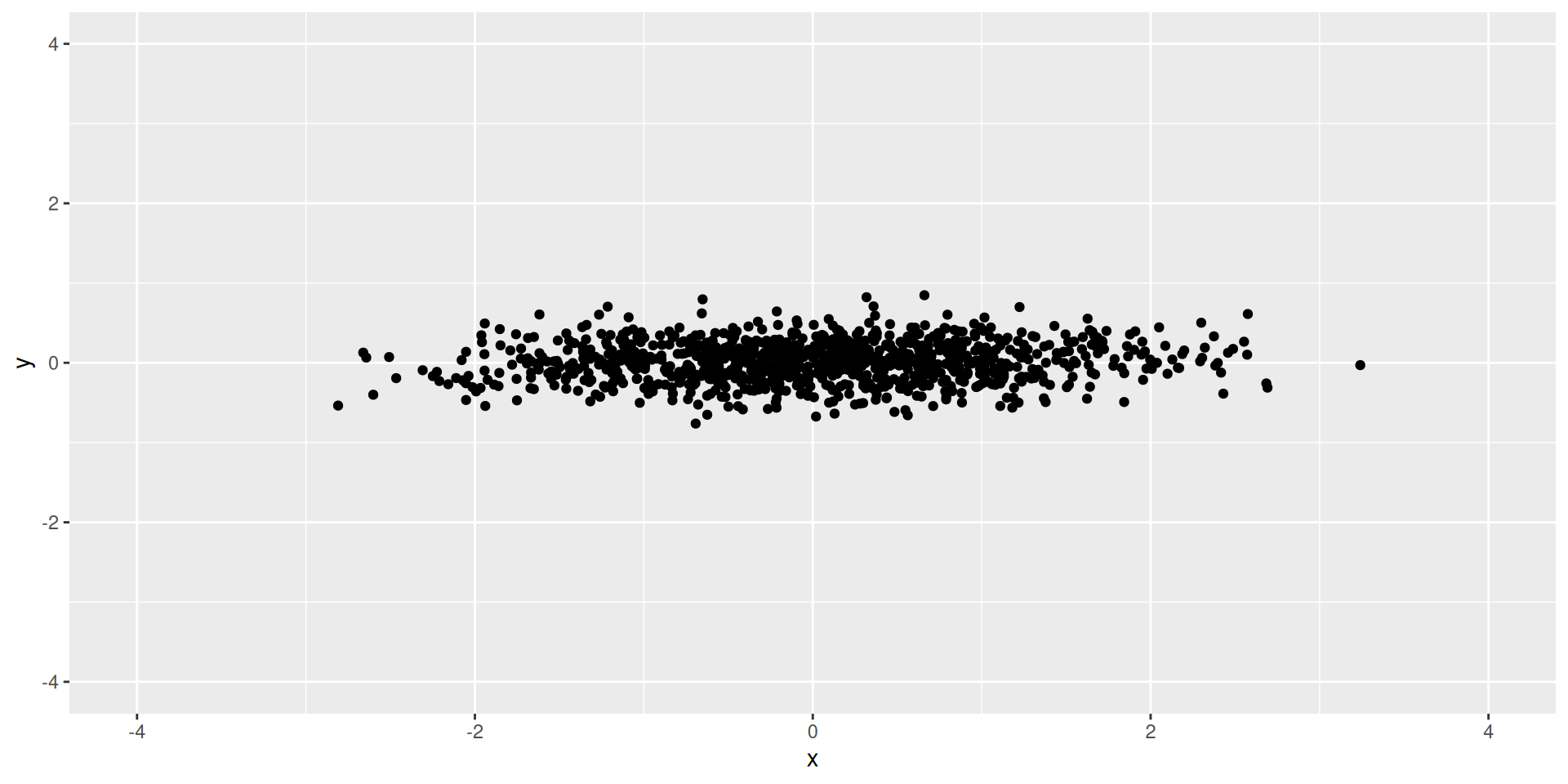
How much variance is explained by each of the PC’s?
What proportion of variance is explained by each of the PC’s?
| name | value | proportion |
|---|---|---|
| var1 | 0.9834589 | 0.9391552 |
| var2 | 0.0637151 | 0.0608448 |
- 93% of our variance (information) is contained in our first PC
Harder Example
- Exercise: What should the first and second PCs be?
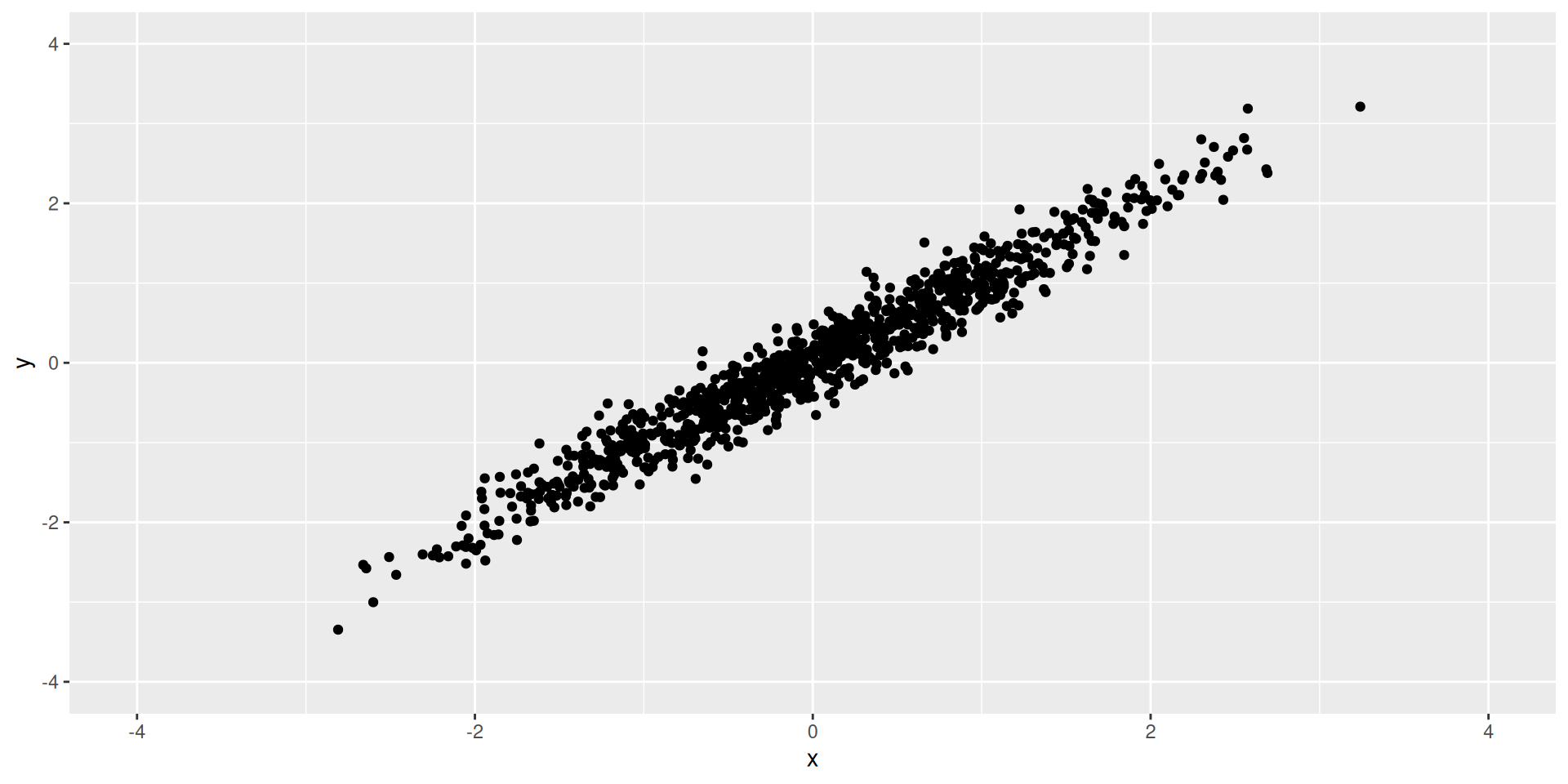
How much variance is explained by each of the PC’s?
Principal Component Analysis
Intuition
- PCA finds new axes (Principal Components) that capture the maximum variance in the data.
- PC1: Direction of maximum variance.
- PC2: Orthogonal to PC1, next maximum variance.
- …
- We can project our data onto these 2-3 new axes to visualize it!
PCA with Recipes
step_pca(): Computes principal components.- Pre-requisites:
- Numeric only: Use
step_dummy()for categorical. - Scaled: Use
step_normalize()(Variance depends on scale!).
- Numeric only: Use
Example: Palmer Penguins
- Let’s visualize the 4 numeric body measurements in 2 dimensions.
Code
library(tidymodels)
library(palmerpenguins)
library(tidyverse)
# Define Recipe
pca_rec <- recipe(~ ., data = penguins) |>
step_rm(year, sex, island) |> # Remove non-measurements
step_naomit(all_predictors()) |>
step_normalize(all_numeric_predictors()) |>
step_pca(all_numeric_predictors()) # Keep top 2 PCs
# Prep and Bake
pca_prep <- prep(pca_rec)
pca_data <- bake(pca_prep, new_data = NULL)
head(pca_data)# A tibble: 6 × 5
species PC1 PC2 PC3 PC4
<fct> <dbl> <dbl> <dbl> <dbl>
1 Adelie -1.84 -0.0476 0.232 0.523
2 Adelie -1.30 0.428 0.0295 0.402
3 Adelie -1.37 0.154 -0.198 -0.527
4 Adelie -1.88 0.00205 0.618 -0.478
5 Adelie -1.91 -0.828 0.686 -0.207
6 Adelie -1.76 0.351 -0.0276 0.504Visualizing PCA
Scatterplot of PC1 vs PC2
- The most common PCA plot.
- Points close together are “similar” in the original high-dimensional space.
Code
# Since we removed 'species' in the recipe to avoid using it in PCA,
# we might want to bind it back for plotting coloring.
# A better workflow is to keep it as an ID/Role but not utilize it in step_pca.
pca_rec_2 <- recipe(species ~ ., data = penguins) |>
step_rm(year, sex, island) |>
step_naomit(all_predictors()) |>
step_normalize(all_numeric_predictors()) |>
step_pca(all_numeric_predictors(), num_comp = 2)
# Default behavior: step_pca ignores 'outcome' variables, which is handy!
pca_prep_2 <- prep(pca_rec_2)
pca_juice <- juice(pca_prep_2) # juice() is a shortcut for bake(prep, new_data=NULL)
ggplot(pca_juice, aes(x = PC1, y = PC2, color = species)) +
geom_point(size = 3, alpha = 0.8) +
theme_minimal() +
labs(title = "Penguins PCA", subtitle = "Species separate well in PC space")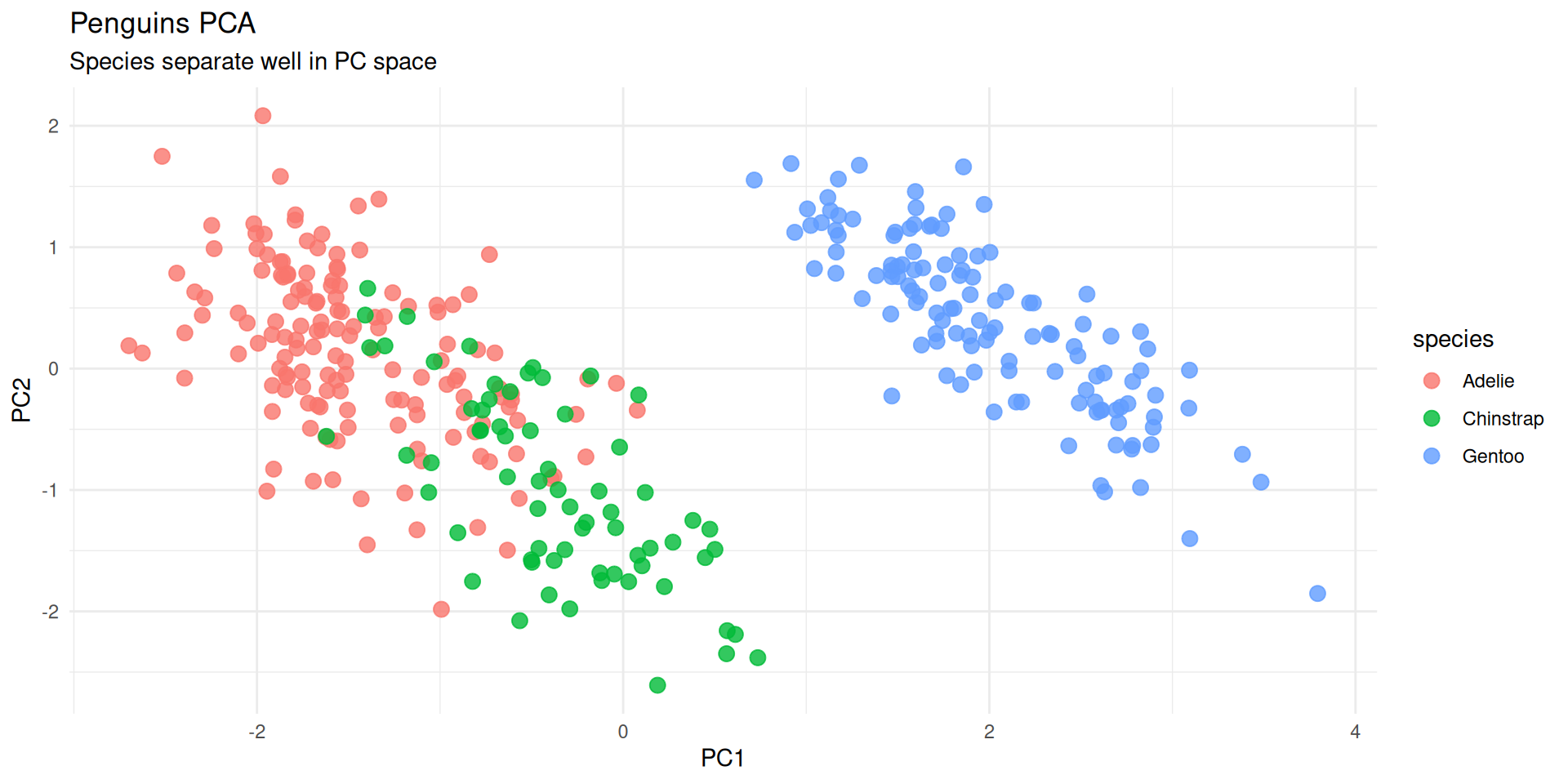
Loadings: What do the PCs mean?
- Loadings: The contribution of each original variable to the PC.
- We can extract them from the prepped recipe using
tidy().
Code
pca_comps <- tidy(pca_prep_2, number = 2, type = "coef") # number=2 refers to the step number index if unknown, typically easier to look up.
# Better way: ID the step
pca_rec_named <- recipe(species ~ ., data = penguins) |>
step_naomit(all_predictors()) |>
step_normalize(all_numeric_predictors()) |>
step_pca(all_numeric_predictors(), id = "pca")
pca_prep_named <- prep(pca_rec_named)
pca_comps <- tidy(pca_prep_named, id = "pca")
pca_comps |>
filter(component %in% c("PC1", "PC2")) |>
ggplot(aes(x = value, y = terms, fill = terms)) +
geom_col() +
facet_wrap(~component) +
theme_minimal() +
theme(legend.position = "none") +
labs(title = "PCA Loadings: Contribution of variables")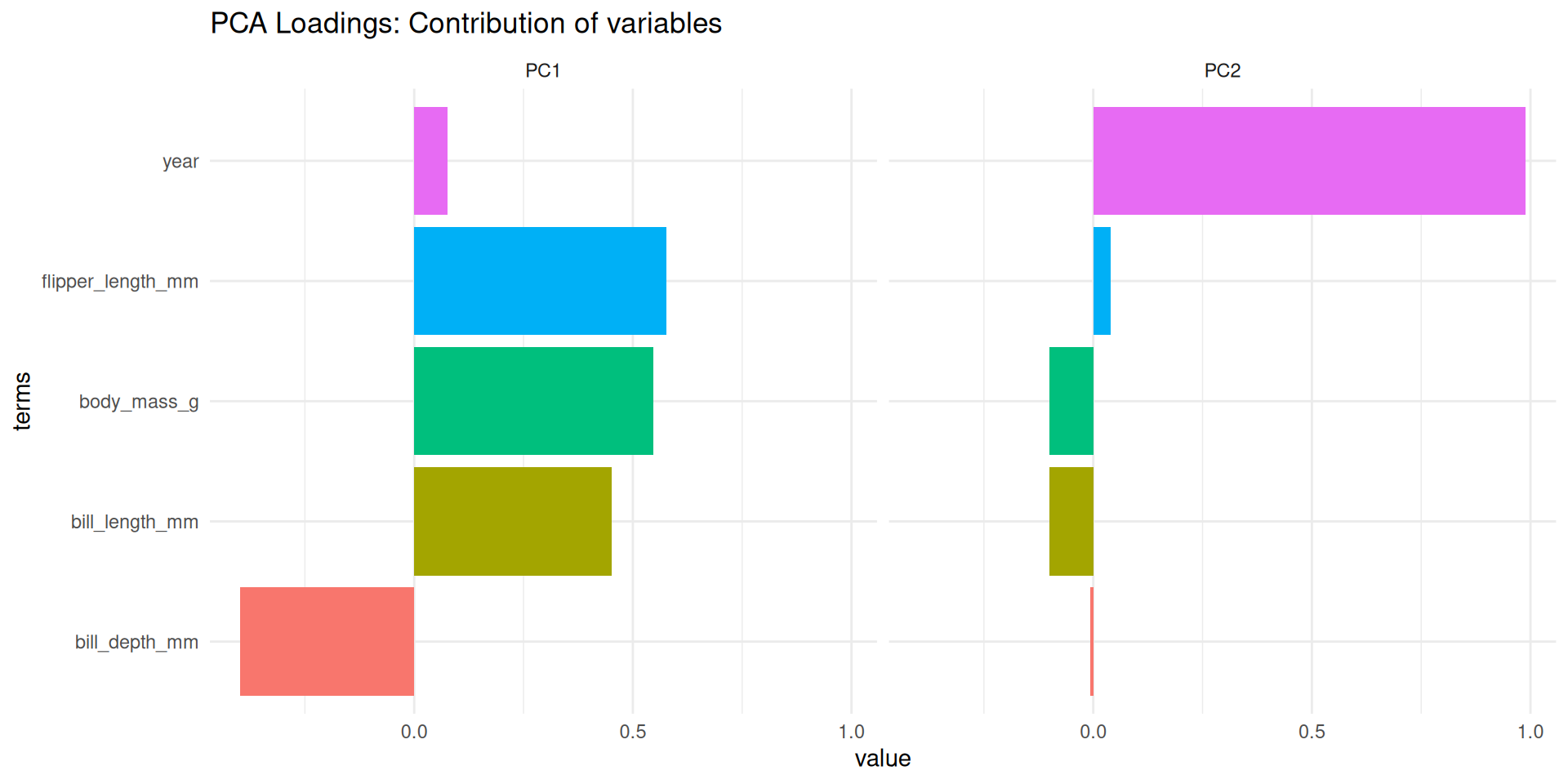
Scree Plot: How many PCs?
- How much variance is explained by each component?
Code
# Extract variance explained
pca_var <- tidy(pca_prep_named, id = "pca", type = "variance")
pca_var |>
filter(terms == "percent variance") |>
ggplot(aes(x = component, y = value)) +
geom_col(fill = "steelblue") +
geom_line(group = 1) +
geom_point() +
labs(title = "Scree Plot", y = "% Variance Explained") +
theme_minimal()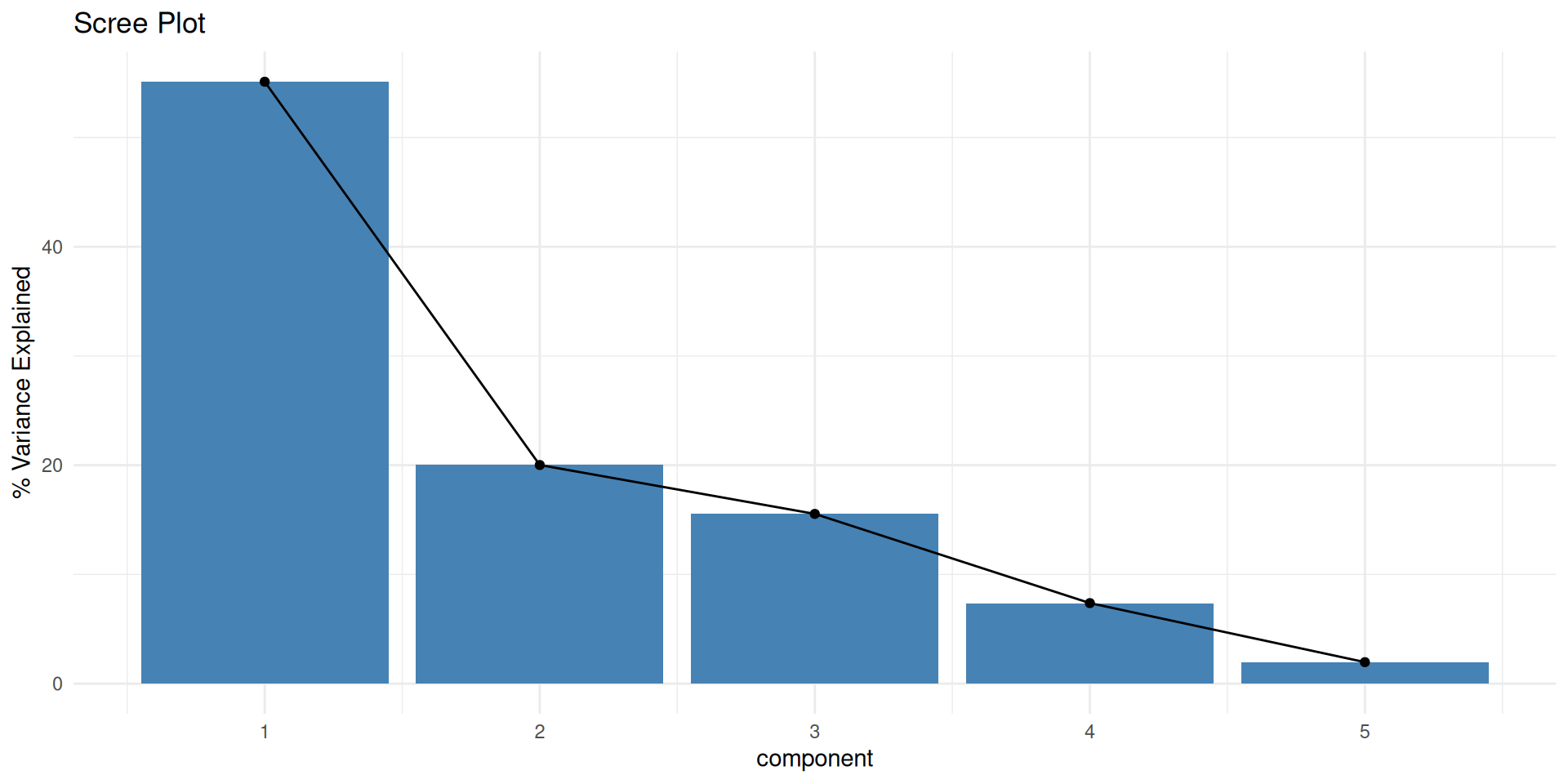
Wrap-Up
Recap
- Dimensionality Reduction: Visualizing high-D data in 2D/3D.
- Steps:
step_normalize()(Crucial!)step_pca()
- Visualization:
- Score Plot: PC1 vs PC2 (Where are the data points?)
- Loadings Plot: Terms contribution (What do the axes mean?)
- Scree Plot: Variance explained (How much info is kept?)
Do Next
- Read Chapter 16: Dimensionality Reduction from Tidy Modeling with R.
- There’s NO recitation Gem for this textbook but I recommend creating your own and adding the textbook chapter and these slides.
- That’s it for tonight!
